Entry information : LuPrx192 (Lus10035925)
| Entry ID | 12128 |
|---|---|
| Creation | 2013-05-28 (Qiang Li) |
| Last sequence changes | 2014-05-23 (Christophe Dunand) |
| Sequence status | complete |
| Reviewer | Not yet reviewed |
| Last annotation changes | 2014-05-23 (Christophe Dunand) |
Peroxidase information: LuPrx192 (Lus10035925)
| Name (synonym) | LuPrx192 (Lus10035925) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Class | Class III peroxidase [Orthogroup: Prx020] | ||||||||||||||||||||||||||||||||||||
| Taxonomy | Eukaryota Viridiplantae Streptophyta Linaceae Linum | ||||||||||||||||||||||||||||||||||||
| Organism | Linum usitatissimum (flax) [TaxId: 4006 ] | ||||||||||||||||||||||||||||||||||||
| Cellular localisation | N/D |
||||||||||||||||||||||||||||||||||||
| Tissue type | N/D |
||||||||||||||||||||||||||||||||||||
| Inducer | N/D |
||||||||||||||||||||||||||||||||||||
| Repressor | N/D |
||||||||||||||||||||||||||||||||||||
| Best BLASTp hits |
|
||||||||||||||||||||||||||||||||||||
| Gene structure Fichier |
Exons►
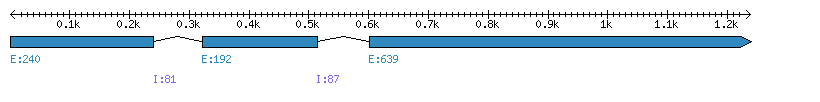
|
||||||||||||||||||||||||||||||||||||
Literature and cross-references LuPrx192 (Lus10035925)
| Literature | Day A, Fénart S, Neutelings G, Hawkins S, Rolando C, Tokarski C. Identification of cell wall proteins in the flax (Linum usitatissimum) stem. Proteomics. 2013 13(5):812-25. |
|---|---|
| DNA ref. | GenBank: AFSQ01019756.1 Phytozome 12: scaffold76 (290807..291826) |
| EST ref. | GenBank: JG237679.1 (28363..29601) [3' end] |
Protein sequence: LuPrx192 (Lus10035925)
| Sequence Properties first value : protein second value (mature protein) |
|
||||||||||||
| Sequence Send to BLAST Send to Peroxiscan |
|
||||||||||||
| Remarks | Complete sequence from genomic. Incorrect prediction from Phytozome (exon 1 is missing but not found in the genomic, longer 3' found from ESTs => VANGGRRQVSNRLSSFS). Experimental evidency of cell wall localization. | ||||||||||||
| DNA ► Send to BLAST |
|
||||||||||||
| CDS► Send to BLAST |
|
||||||||||||
