Entry information : ZmAPx03(Zm00001d025047 / GRMZM2G004211 [6a] / APx3)
| Entry ID | 6379 |
|---|---|
| Creation | 2008-10-03 (Christophe Dunand) |
| Last sequence changes | 2011-01-21 (Christophe Dunand) |
| Sequence status | complete |
| Reviewer | Christophe Dunand |
| Last annotation changes | 2023-01-03 (Christophe Dunand) |
Peroxidase information: ZmAPx03(Zm00001d025047 / GRMZM2G004211 [6a] / APx3)
| Name | ZmAPx03(Zm00001d025047 / GRMZM2G004211 [6a] / APx3) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Class | Ascorbate peroxidase [Orthogroup: APx2005] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy | Eukaryota Viridiplantae Streptophyta Monocotyledons Poaceae Zea | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Organism | Zea mays [TaxId: 4577 ] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular localisation | N/D |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue types | Meristems Ovaries Roots |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Inducer | N/D |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Repressor | N/D |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Best BLASTp hits |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene structure Fichier |
Exons►
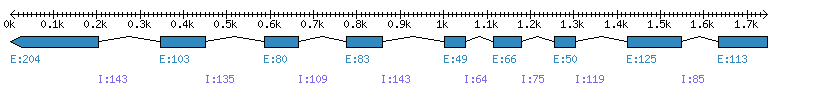
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Literature and cross-references ZmAPx03(Zm00001d025047 / GRMZM2G004211 [6a] / APx3)
| Protein ref. | UniProtKB: B6T684 |
|---|---|
| EST ref. | GenBank: DR972292.1 [5' end] CF625763.1 [3' end] |
| Cluster/Prediction ref. | UniGene: Zm.28302 |
Protein sequence: ZmAPx03(Zm00001d025047 / GRMZM2G004211 [6a] / APx3)
| Sequence Properties first value : protein second value (mature protein) |
|
||||||||||||
| Sequence Send to BLAST Send to Peroxiscan |
|
||||||||||||
| Remarks | Complete sequence from genomic (chromo 10, 9 introns) and 6 ESTs. Incorrect prediction from Phytozome due to a probable sequencing error. ACC CCC CC(C)T CGC TCT, (c) has been removed from the genomic sequence to remove the frame shift. | ||||||||||||
| DNA ► Send to BLAST |
|
||||||||||||
| CDS► Send to BLAST |
|
||||||||||||
