Entry information : EgrPrx05(Eucgr.A00043 / Egrandis_v1_0.043198m)
| Entry ID | 8105 |
|---|---|
| Creation | 2011-02-04 (Christophe Dunand) |
| Last sequence changes | 2016-03-21 (Catherine ) |
| Sequence status | complete |
| Reviewer | Not yet reviewed |
| Last annotation changes | 2022-03-28 (Catherine ) |
Peroxidase information: EgrPrx05(Eucgr.A00043 / Egrandis_v1_0.043198m)
| Name | EgrPrx05(Eucgr.A00043 / Egrandis_v1_0.043198m) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Class | Class III peroxidase [Orthogroup: Prx080] | ||||||||||||||||||||||||||||||||||||
| Taxonomy | Eukaryota Viridiplantae Streptophyta Myrtaceae Eucalyptus | ||||||||||||||||||||||||||||||||||||
| Organism | Eucalyptus grandis [TaxId: 71139 ] | ||||||||||||||||||||||||||||||||||||
| Cellular localisation | N/D |
||||||||||||||||||||||||||||||||||||
| Tissue type | N/D |
||||||||||||||||||||||||||||||||||||
| Inducer | N/D |
||||||||||||||||||||||||||||||||||||
| Repressor | N/D |
||||||||||||||||||||||||||||||||||||
| Best BLASTp hits |
|
||||||||||||||||||||||||||||||||||||
| Gene structure Fichier |
Exons►
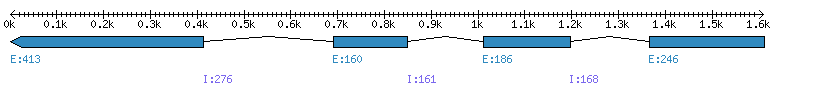
|
||||||||||||||||||||||||||||||||||||
Literature and cross-references EgrPrx05(Eucgr.A00043 / Egrandis_v1_0.043198m)
| DNA ref. | Phytozome 12: scaffold_1 (423824..422215) |
|---|---|
| Cluster/Prediction ref. | Genebank: 104450810 |
Protein sequence: EgrPrx05(Eucgr.A00043 / Egrandis_v1_0.043198m)
| Sequence Properties first value : protein second value (mature protein) |
|
||||||||||||
| Sequence Send to BLAST Send to Peroxiscan |
|
||||||||||||
| Remarks | Complete sequence from genomic (3 introns). Incorrect prediction from Phytozome (A short sequence in frame is missing). No EST. | ||||||||||||
| DNA ► Send to BLAST |
|
||||||||||||
| CDS► Send to BLAST |
|
||||||||||||
