Entry information : TsPrx[P]49-1(Thhalv10000286m / ThPrx49-1)
| Entry ID | 9781 |
|---|---|
| Creation | 2011-11-18 (Qiang Li) |
| Last sequence changes | 2011-11-28 (Qiang Li) |
| Sequence status | theoretical translation / pseudogene |
| Reviewer | Christophe Dunand |
| Last annotation changes | 2023-06-15 (Christophe Dunand) |
Peroxidase information: TsPrx[P]49-1(Thhalv10000286m / ThPrx49-1)
| Name | TsPrx[P]49-1(Thhalv10000286m / ThPrx49-1) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Class | Class III peroxidase [Orthogroup: Prx004]* | ||||||||||||||||||||||||||||||||||||
| Taxonomy | Eukaryota Viridiplantae Streptophyta Brassicaceae Eutrema | ||||||||||||||||||||||||||||||||||||
| Organism | Thellungiella salsuginea (Eutrema salsugineum) [TaxId: 72664 ] | ||||||||||||||||||||||||||||||||||||
| Cellular localisation | N/D |
||||||||||||||||||||||||||||||||||||
| Tissue type | N/D |
||||||||||||||||||||||||||||||||||||
| Inducer | N/D |
||||||||||||||||||||||||||||||||||||
| Repressor | N/D |
||||||||||||||||||||||||||||||||||||
| Best BLASTp hits |
|
||||||||||||||||||||||||||||||||||||
| Gene structure Fichier |
Exons►
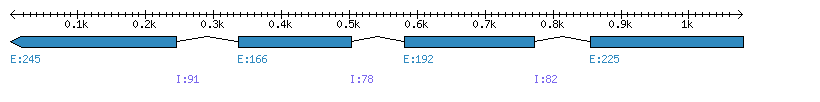
|
||||||||||||||||||||||||||||||||||||
Literature and cross-references TsPrx[P]49-1(Thhalv10000286m / ThPrx49-1)
| DNA ref. | Phytozome 12: scaffold_15 (2882568..2881490) |
|---|
Protein sequence: TsPrx[P]49-1(Thhalv10000286m / ThPrx49-1)
| Sequence Properties first value : protein second value (mature protein) |
|
||||||||||||
| Sequence Send to BLAST Send to Peroxiscan |
|
||||||||||||
| Remarks | Partial sequence from genomic. Incorrect prediction from phytozome (3' end is missing which was not found yet). No EST. | ||||||||||||
| DNA ► Send to BLAST |
|
||||||||||||
| CDS► Send to BLAST |
|
||||||||||||
