A. What's GECA?
GECA (Gene Evolution/Conservation Analysis) is a collection of perl scripts, which align exon/intron structures and detect common introns and similarities between sequences in order to provide information about genes evolution and/or conservation.
A local version of GECA can be installed on UNIX platforms and requires pre-installation of PERL, MAFFT and CIWOG.The strategy relies on a simple fact : by aligning the commun introns of closely related sequences, we align the exon structure of the respective genes. Once the sequences are aligned, the are compared, amino acide by amino acide, to search for similarity between the sequences.
A web based version is available at "https://peroxibase.toulouse.inra.fr/geca/".
Comments and questions are welcome.B. Input format
The data submitted by the user should be in FASTA format. The FASTA header is as follows :">Accession_id|sequence name" for exemple : 5546|Sb1CysPrx01An example of the protein sequences format:
1064|OsPrx54
MALLLLRRGGGFAAATVLAVVVVALVLSCGGGAEAAVRDLRVGYYAETCPDAEAVVRDTMARARAHEARSVASVMRLQFH
>1049|OsPrx39
MAATLRWGGGGLAVAAFAAVVALSGLLGVAANYGGGGGFLFPQFYQHTCPQMEAVVGGIVARAHAEDPRMAASLLRMHFH An example of the genomic sequences format:
>1064|OsPrx54
ATGGCGGCGACATTGCGTTGGGGCGGCGGCGGGCTCGCGGTGGCGGCGTTTGCGGCGGTGGTCGCGTTGTCCGGCCT
CCT
>1049|OsPrx39
ATGGGCGCTGTGGCTGCGGTTCGTGCCGCGGTCCTGGTCGTGGCCGTGGCCCTCGCCGCGGCGGCGGCCGGCGCGTC
GGC The gene structure information given in Genbank format should be preceded by the same FASTA header. For exemple :
>1064|OsPrx54
join(203122798..203122892,203126718..203126874,203130660..203130806,203131568..203131714,
203133072..203133200)
>1049|OsPrx39
complement(join(1..672))C. GECA Results:
Once the data is submitted, you will be directed to the GECA Results page. This page gives access to the different results generated by GECA, which are: *the multiple alignment file, *CIWOG Results, sequences of the common introns detected and the image generated by GECA.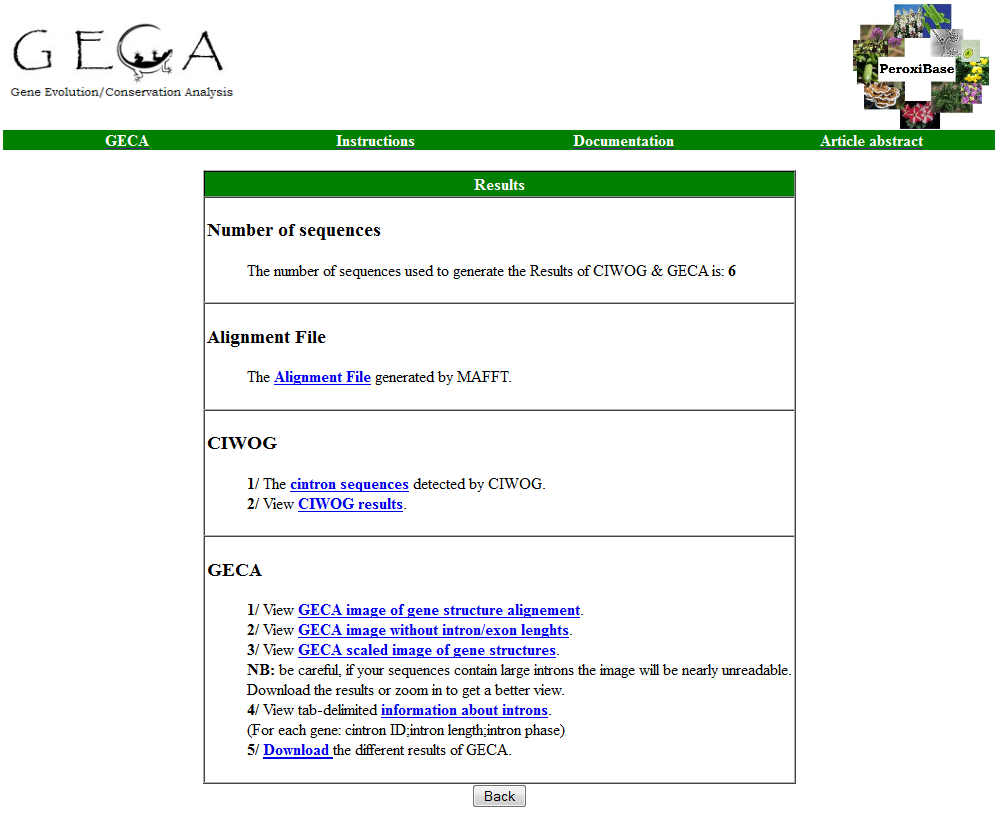
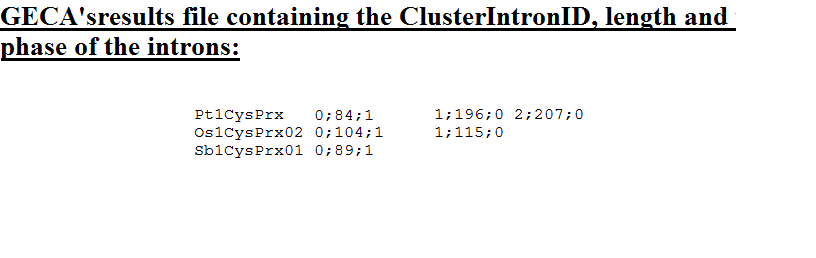

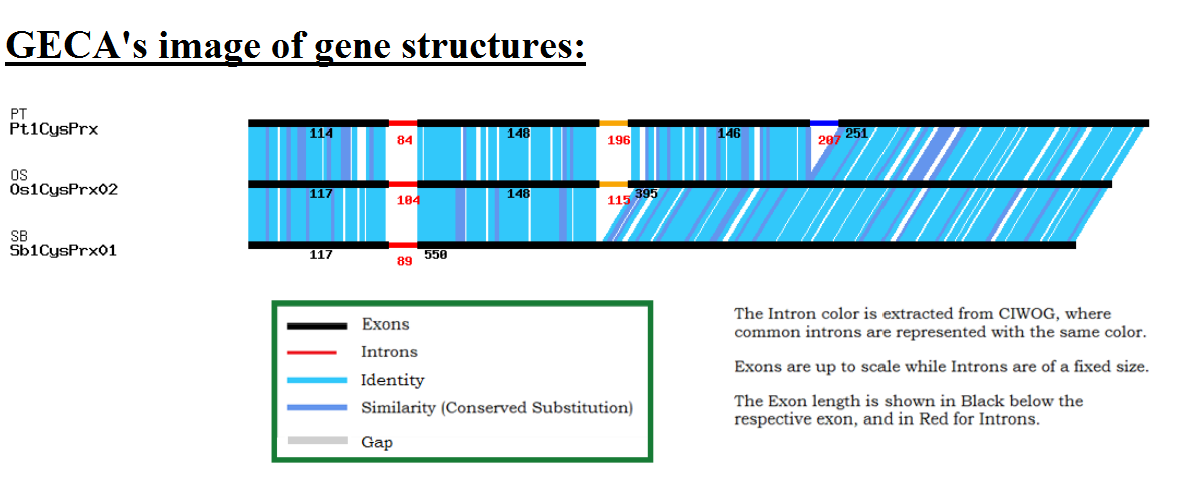
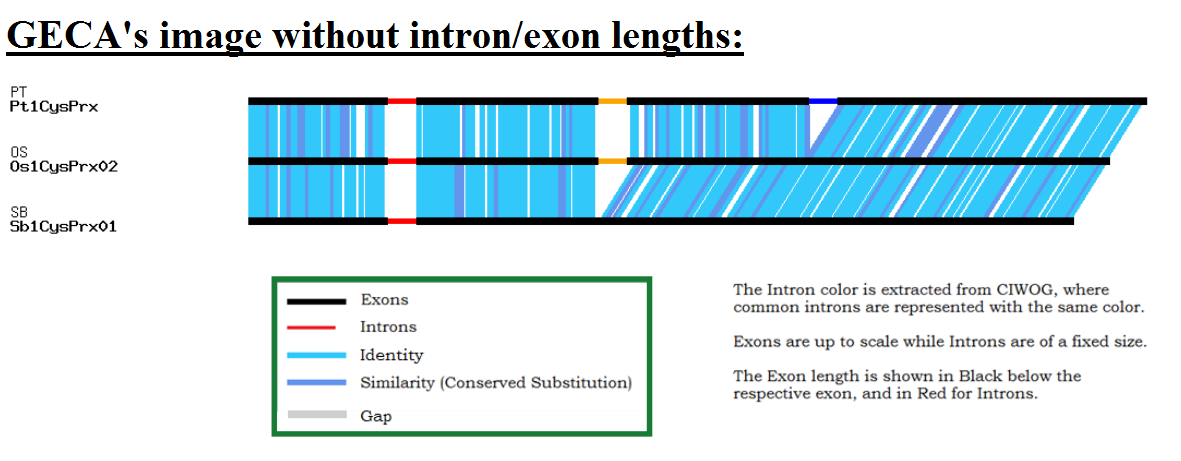
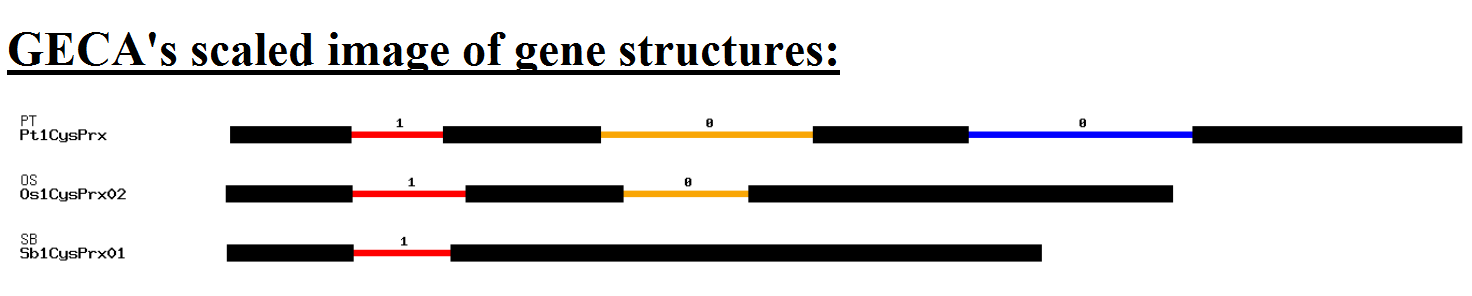

Authors and Help
GECA has been written by Nizar Fawal (UMR 5546 CNRS/Universite Paul Sabatier).
For Questions, please write to : Peroxibase@lrsv.ups-tlse.fr GECA home page is at "https://peroxibase.toulouse.inra.fr/"Citation
Please use the following article when citing GECA:
Fawal, N., Savelli, B., Dunand, C., Mathé, C., GECA : a fast tool for Gene Evolution and Conservation Analysis in eukaryotic protein families.
Bioinformatics. Application Notes. 2012. PMID:22467908.
